Botany online 1996-2004. No further update, only historical document of botanical science!
In our discussion of photosynthesis have we thus far only regarded biochemical reactions. In the section about glycolysis and other biosynthetic pathways was the significance of single enzymes pointed out. Since quite some time know, for example, have all enzymes involved in glycolysis been purified and isolated and each step of the pathway can be analyzed under in vitro conditions. It has also been tried to isolate the complete set of components necessary for photosynthesis in order to reconstitute the whole system. But all attempts failed because the premises they were based on were wrong as we know today.
A number of problems have not been taken into consideration until now. The terms photosystem I and photosystem II, for example, have been introduced and all participating pigments have been mentioned but the following subjects remain to be discussed:
![]() How are the photosystems organized?
How are the photosystems organized?
![]() How are the pigments arranged?
How are the pigments arranged?
![]() Why does one of the chlorophyll molecules react different
than all the others?
Why does one of the chlorophyll molecules react different
than all the others?
![]() Why are action and absorption spectra not quite congruent?
Why are action and absorption spectra not quite congruent?
![]() Why reacts P 680 (chlorophyll a) different than P 700 (chlorophyll
a, too)?
Why reacts P 680 (chlorophyll a) different than P 700 (chlorophyll
a, too)?
![]() How are electron transport chain and ATP production coupled?
How are electron transport chain and ATP production coupled?
![]() How are photosystem I and II linked?
How are photosystem I and II linked?
![]() Which structural prerequisites have to exist in order for
the two systems to co-operate?
Which structural prerequisites have to exist in order for
the two systems to co-operate?
It was always accepted that each of the biochemical reactions was catalyzed by a specific enzyme and still, it took quite some time before it was realized that the chlorophyll and the other pigments are protein-bound and that they are only active as protein-chlorophyll (and protein-pigment, respectively) complexes. The isolated pigments themselves were useless for photosynthesis. The pigment-protein complex, (most) proteins of the electron transport chain as well as the catalyst of ATP synthesis (ATP synthase) are integral compounds of the photosynthesis membrane(s) (= the thylacoid membranes of algae and higher green plants, cytoplasmatic membranes of photosynthetically active bacteria and blue-green algae). The location within the membrane (at the out- or the inside, for example) and the relative arrangement of the proteins towards each other are important prerequisites of energy transformation.
This is not only true for photosynthetic reactions but also for those of the respiratory chain and for the enzymes located within the purple membrane of Halobacterium halobium (an archaebacterium using light energy for the production of ATP without an electron flow).
The requirements for energy transformation are even higher: completely intact membranes that are impermeable for protons and that enclose compartments thus maintaining a stable electrochemical gradient between inside and outside. The production of ATP is based on a directed proton dislocation paralleled by a change of the compartment's pH and of its membrane potential.
The research into the proteins essential for photosynthesis started very late. The reason is that all of them are membrane-bound which rendered it nearly impossible to isolate and characterize them with the classical methods of protein analysis.
Only after sensitive techniques like gel electrophoresis and the controlled use of detergents like sodium dodecyl sulfate (SDS) had been developed, became it possible to separate the proteins and to identify them as bands in a gel. A side product of this technique is the determination of the molecular weights of the respective polypeptide chain.
A second, independent attempt was and is the use of specific probes like fluorescence-tagged antibodies that help to find out whether a certain protein (or part of a polypeptide chain) is located at the inside or the outside of a membrane. The use of antibodies against specific proteins allows, too, to precipitate these proteins selectively since only they are able to form the extremely specific antigen - antibody complex.
Cross-linking agents render it possible to elucidate the surrounding of a molecule. And the use of specific inhibitors helps localizing their site of effect. DCMU [3-(3', 4' - dichlorphenyl) - 1,1 - dimethylurea] has since years been used to inhibit photosystem II. It has no effect on photosystem I and was therefore used by ARNON and his collaborators as an important help to study the electron transport chain that starts at photosystem I independently of that induced by photosystem II.
We know today that DCMU does not effect chlorophyll itself but a certain protein, the plastoquinone-binding protein.
A third possibility to characterize the photosynthetic membrane is the analysis of certain mutants. The single-celled alga Chlamydomonas reinhardii proved to be a good test object. Quite a range of mutants with photosynthetic defects are known. They can be grouped in four classes:
- mutants with a defect in photosystem I,
- mutants with a defect in photosystem II,
- mutants with a defect in photophosphorylation and
- mutants with a defect in the antenna complex.
It is quite striking that almost all mutants are characterized not only by the loss or change of a certain polypeptide chain but by the lack of a whole complex, for example that of PS I. It seems therefore as if the mutations would lead to pleiotropic effects. Or, expressed differently: when a polypeptide chain is changed or missing does the assembly of the other polypeptide chains not work any more. This observation shows how tight the interactions between the single polypeptide chains are and how important they are for their mutual co-operation.
A further and not less important technique is electron microscopy usually used in combination with freeze-etching.
The sequencing of membrane proteins remains difficult. And yet, the sequences of most proteins involved in photosynthesis could be determined during the last years via the sequencing of their respective genes. The most remarkable outcome of this work is that these proteins contain (just like proteins of animal or bacterial membranes, too) a large portion of alpha - helices. The lengths of the helices corresponds to the thickness of the membranes. The single helices are connected via polar and / or non-hydrophobic sequences.
Several chlorophyll-binding proteins of the photosynthetic membranes of different systematic groups (angiosperms, gymnosperms, algae, bacteria) have been isolated and characterized. The best-known are P 700-chlorophyll-a-protein 1 and the light-harvesting chlorophyll-a/b-protein 2 which have been studied in the laboratory of J. P. THORNBERGER at the University of California, Los Angeles. Both are strongly hydrophobic, integral membrane proteins. Both bind chlorophyll a but only the latter binds chlorophyll b, too. The P 700-chlorophyll-a-protein 1 contains the reaction centre (P 700) of photosystem I, i.e. one of the chlorophyll molecules is bound in a specific configuration and is located in a surrounding (due to a specific amino acid composition and the folding of the polypeptide chain) that renders it different than all other chlorophyll molecules bound by this protein, too. This structural peculiarity is the precondition for the light-induced activation and consequently for the induction of the electron flow.
The molecular weight of the polypeptide chain is 110,000 Dalton. It is able to bind 14 chlorophyll molecules. The light-harvesting-protein (light-harvesting chlorophyll-a/b-protein 2) is also very common. It is mainly associated with photosystem II but effects on photosystem I have been observed, too. Chlorophyll a and b are bound in equimolar amounts besides lutein and beta - carotene. The chlorophyll to carotenoid ratio is 3 - 7 : 1 on a molecular level.
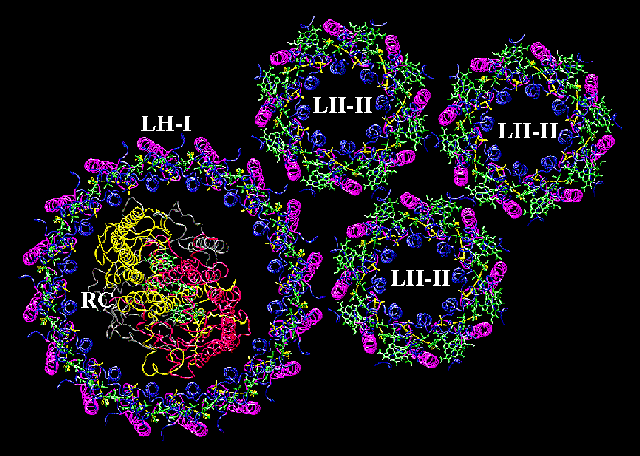
©: Theoretical Biology Group - University of Illinois at Urbana-Champaign
ATP - synthase is the enzyme that catalyzes the synthesis of ATP. Since the production of ATP occurs not only during photosynthesis but during respiration, too, suggested the idea that the ATP production of both cases is based on similar mechanisms itself.
After having collected experience with the mitochondrial ATP synthase, did E. RACKER of the Cornell University isolate an enzyme of thylacoid membranes that resembled the respective mitochondrial enzyme very much. In the electron microscope did it look like a stemmed knob. The knob was termed CF1 and the stem CF0 (F1 and F0 respectively in the mitochondrial enzyme). CF0 (or F0) is a membrane anchor. While the F1 - F0 complex is localized in the inner mitochondrial membrane and the knob is directed towards the mitochondrial matrix are the respective molecular parts of the CF0 - CF1 complex found at the outside of the thylacoid membranes. In both cases consists the ATP synthase out of several different polypeptide chains, it is an enzyme complex. The phosphorylation of ADP works only, if the ATP synthase is a component of an intact, proton-permeable membrane. It has to separate two compartments (the inner part of the vesicle and the surrounding).
Since quite some time has the ATP synthase been ascribed the function of a coupling factor. This means that it is able to utilize the free energy released by electron transport. Such energy conservation is referred to as energy coupling or energy transduction.
How does this work? It might have been assumed that the electron transport chain serves the production of energy-rich intermediates and that these constitute an energy store for the production of ATP. Two arguments against this idea exist:
- No such substances have ever been isolated and
photophosphorylation (and the oxidative phosphorylation of the respiratory chain, respectively) is only possible if the thylacoid membranes (the inner mitochondrial membrane, respectively) are intact.
Another assumption had been that the ATP synthase changes its configuration and is thus itself transferred into an activated state. In such a case would the energy be transiently stored in weak interactions. This hypothesis, too, failed to withstand experimental scrutiny.
In 1961 proposed P. MITCHELL (Glynn Research Laboratories, Great Britain) that the energy set free during the electron transport is conserved as a proton gradient across the membrane. The energy would then not be stored as a chemical bond but as an electrochemical gradient. The electrochemical potential of this gradient would be harnessed to synthesize ATP. The hypothesis explains several key observations:
A very convincing experiment was performed by A. T. JAGENDORF (1966, Cornell University, Ithaca, N. Y.): he took isolated thylacoids and incubated them in a pH buffer until the same pH (4) was measured at both sides of the membrane. After the equilibrium had been achieved, did he quickly transfer the thylacoids to a media of pH 8 containing both ADP and Pi. Immediately after the transfer was the pH between inside and outside evened out while, at the same time, the system produced ATP. The proton flow across the membrane was used for the production of ATP. The experiment worked only with intact membranes. In addition have all biochemical processes to be directed (vectorial). All enzyme molecules have to have the same direction so that the protons are transported in only one direction.
- It is consistent with the fact that oxidative phosphorylation requires an intact inner mitochondrial membrane.
- The inner membrane is impermeable to ions like H+ or OH- whose free diffusion would discharge the electrochemical gradient.
- The electron transport results in the transport out of intact mitochondria (out of the thylacoid space of chloroplasts) thereby creating a measurable electrochemical gradient across the inner mitochondrial membrane (the thylacoid membrane of chloroplasts).
- Substances that increase the permeability of the inner mitochondrial or the thylacoid membrane to protons, thereby dissipating the electrochemical gradient, allow electron transport to continue but inhibit ATP synthesis: they 'uncouple' electron transport from oxidative phosphorylation.
How can a flow of protons induce the production of ATP??
To understand the process is it useful to study the ATP production a little more. ADP and phosphate (Pi) are its starting compounds. It is known that both are bound to neighbouring but separate binding sites of the enzyme complex (the ATP synthase ). To produce ATP (ADP~P) has one H+ to be removed from ADP and one OH- from phosphate. In a formal sense is water split off. As soon as the ions have left the complex do both combine with their counterions to water (not with each other). The end result is a directed flow of protons.
This does not mean that one proton is transferred through the membrane via the ATP synthase complex. Instead is a newly formed proton given off into solution at one side while another proton is captured and neutralized (by a OH- ion) at the other side of the membrane.
Independent of these observations could GRÄBER and WITT (Max-Vollmer-Institut, Technische Universität Berlin) show that a direct coupling between proton gradient and the electron transport chains of the photosystems I and II exists. It emerged that the already known 'Z'-scheme is no hypothetical product but that the involved components are arranged like a Z within the photosynthetic membrane, i.e. the structural basis for the process discussed in the previous sections became known.
D. von WETTSTEIN and R. P. OLIVER (Carlsberg Laboratorium, Copenhagen, 1985) summarized all results and were thus able to develop a model that explains the topology of the single protein complexes.
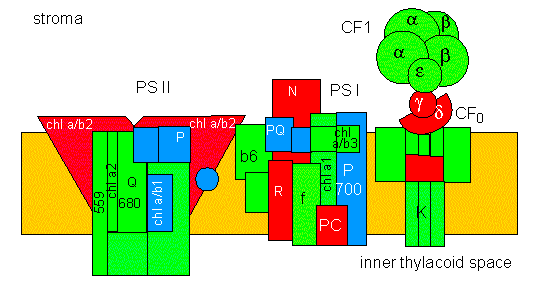
The photosynthetic membrane contains four complexes altogether. Each has subunits encoded in the nucleus and others encoded by plastids. Several proteins bind chlorophyll a, one binds chlorophyll a and b. The PS II complex is mainly localized in stacked, PS I and the ATP synthase complex (CF1 - CF0) in non-stacked thylacoid membranes.
A further remark: the reactions of the CALVIN cycle are catalyzed by soluble enzymes, localized within the stroma.
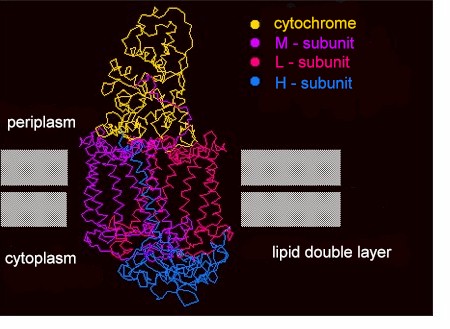
In 1985 was the structure of the photosynthetic reaction centre's protein subunits determined. J. DEISENHOFER, H. MICHEL and R. HUBER (Max-Planck-Institut für Biochemie, Martinsried) succeeded in crystallizing the protein complex of the photosynthetic membrane of the bacterium Rhodopseudomonas viridis, they determined the folding of the polypeptide chain and the arrangement of the chlorophyll molecules. In 1988 were they awarded the Nobel prize for this work. Together with the tertiary and quaternary structure was also the amino acid sequence of the involved polypeptides determined. The central part of the complex contains two subunits, L and M, each of which forms 5 helices that span the photosynthetic membrane. Two further polypeptides, H and a cytochrome c - like protein are associated. Furthermore belong 4 covalently linked heme groups, 4 bacteriochlorophyll b molecules, 2 molecules of bacteriopheophytin b, 2 quinones, 1 iron ion that is not linked to a heme group as well as carotenoids as prosthetic groups to the complex. The structures found in bacteria are homologous to the reaction centres of the photosystems I and II of green plants.
© Peter v. Sengbusch - Impressum