|
|
Quiz Me! |
Microbial genetics: recombination and plasmids |
|
|
|
Lecture Index | ||
|
|
Course Resources page |
Last revised: Friday, January 7, 2000
Ch. 14 in Prescott et al, Microbiology, 4th Ed.Note: These notes are provided as a guide to topics the instructor hopes to cover during lecture. Actual coverage will always differ somewhat from what is printed here. These notes are not a substitute for the actual lecture!Copyright 2000. Thomas M. Terry
Gene transfer and Recombination
General problem
- Bacteria don't have sex. Normal mechanism of cell division is clonal (one cell produces many identical offspring).
- What possibilities exist for transfer of DNA from one cell to another? How common are they in nature?
- Originally thought that bacteria lacked any significant gene transfer. But discovery of R-plasmid transfer was shocking revelation of rapid gene transfer, not just among individuals of same species but among quite different genera of bacteria.
- Have discovered 3 mechanisms for DNA transfer in bacteria: transformation, transduction, conjugation. Not clear how important these are in nature. In laboratory they are very important.
- Genetic Engineering relies critically on ability to move DNA from one cell to another. Any modern biologist should be familiar with basic mechanisms, possibility of extending these with laboratory "tricks".
Recombination
- Distinguish two stages of gene transfer:
- getting DNA from Donor cell to recipient cell
- getting DNA integrated into recipient (or into a different type of stable form, typically a plasmid).
- Even if 1 happens, no detectable result unless 2 also happens. See figure below:
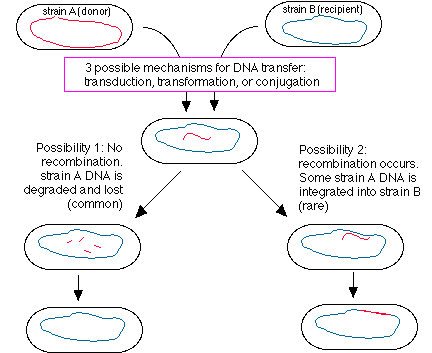
- How does recombination work? Requires specific proteins produced in recipient cell (rec gene products, others). Requires regions of very similar DNA sequence between donor and recipient DNA (homology). Possible for ds DNA to separate, one strand of donor to pair up with one strand of recipient, allow cross-over that attaches donor in place of normal recipient DNA for some distance.
- Why would such a system ever evolve? Probably as a repair mechanism for situations when cell faces double stranded damage to chromosome (can't be fixed by normal repair mechanisms, since don't have a remaining "good strand" to copy from). But if cell had duplicated chromosome but not yet divided, could copy good DNA from other chromosome, use to replace damaged section.
Transformation
- Uptake of DNA fragments from medium surrounding cell. Requires specific proteins in cell membrane, energy.
- Only found naturally in certain cells; e.g. Pneumococcus, Hemophilus, etc. Not found in E. coli.
- Why did this evolve? Maybe as way to scrounge DNA from dead cells, save trouble of having to make nucleotides from scratch.
Competence
- Cells that can take up DNA are said to be competent. Requires induction of several genes
- Typically occurs during exponential growth, shuts off in stationary phase.
Mechanisms for transforming cells without natural competence systems
- Very few bacterial species have evolved highly efficient transformation systems
- In laboratory, can devise techniques to enhance DNA uptake = "artificially induced competence"
- Example: treat E. coli with high Ca++ concentrations, then chill. DNA uptake now occurs (though not as well as with naturally competent bacteria).
- Electroporation: pulsed electric fields produce short-lived membrane pores, allows DNA movement (either in or out of cell). Useful way to move small pieces of DNA and plasmids between cells.
Transduction
Mechanism of generalized transduction
- DNA phages normally replicate in cell, produce many copies of phage DNA, many copies of phage head coat proteins, finally assemble phage heads by packing phage DNA into new phage heads. Attach tail (if present), open cell, release progeny.
- Host DNA often degraded. But occasionally, piece of partially degraded bacterial DNA is correct size to be packed inside phage coat proteins, phage erroneously packs up a "mistake" = transducing phage.
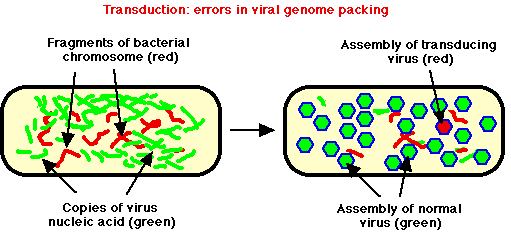
- This "mistake" phage can't cause infection; but it can be transferred to a different bacterium, get DNA into cell without risk of being degraded in environment
- Even if this occurs, chances are slim that successful expression of DNA will occur --- still needs to undergo recombination. If most cells are killed by phage, not much use.
Use of phage P1 as a "gene truck"
- E. coli phage P1 makes this DNA packing mistake about 1 in every 1000 phage particles. Astonishingly high error rate (relative to other phages).
- Can use P1 as very efficient way to move small pieces of DNA from one bacterium (donor) to another (recipient). Maximum size of DNA that can fit in phage head is only about 2% of bacterial chromosome.
- If two genes are close together, can map their relative location by determining cotransduction frequency; e.g., how many times when gene A is transferred by transduction does gene B also get transferred? If frequency is very high, genes are close together. If frequency is zero, they are at least 2% of chromosomal DNA length apart (can't tell more than that, since DNA in phage head is small).
- Generalized transduction = random insertion of any fragment of donor DNA, equal opportunity for transfer to recipient. No preference for any particular piece of DNA being transferred (vs. specialized transduction, a special situation involving only one region of chromosome being transferred -- not responsible for mechanism here)
Conjugation
- Conjugation = plasmid-directed transfer of DNA from one cell to another.
Properties of Plasmids
- Circular DNA elements, always double-stranded DNA, Supercoiled
- Can occur in as few as 1 copy per cell (single copy plasmids) to as many as several dozen (multicopy plasmids).
- Variable sizes; small plasmids about 0.1% size of host chromosome, large plasmids can be as much as 10% the size of host chromosome. Smaller plasmids have few genes (30 or less). Size ranges from 1000 bp (1 kbp) to 1000 kbp.
- Ubiquitous; almost all cells isolated in nature carry plasmids, often more than one kind. (In E. coli alone, more than 300 different plasmids isolated.)
- Have a replicon (origin for DNA replication), number of copies per cell regulated. Large plasmids typically only 1-5 copies/cell (stringent control); small plasmids ~10-50 copies/cell (relaxed control)
- Many plasmids are incompatible; if one is present, cell cannot support another plasmid of same compatibility group.
- Not essential to cell under all circumstances; can be "cured" by agents that impair DNA replication ----> cured cell lacking plasmid. Can be spontaneously lost over time unless some selection makes plasmid valuable to cell.
- Extend range of environments in which a cell can live (e.g., by degrading antibiotics, or providing enzymes for digestion of novel catabolites).
Examples of Plasmid genes
- Antibiotic resistance genes (enzymes that modify or degrade antibiotics) -- plasmids with these genes are called R factors
- Heavy metal resistance (enzymes that detoxify metals by redox reactions)
- Growth on unusual substrates (enzymes for hydrocarbon degradation, etc.)
- Restriction/modification enzymes (protect DNA, degrade unprotected DNA)
- Bacteriocins (proteins toxic to other bacteria lacking the same plasmid)
- Toxins (proteins toxic to other organisms; e.g. humans) -- called virulence plasmids. Some Examples:
- Staph aureus virulence factors: coagulase, hemolysin, enterotoxin, others
- pathogenic E. coli strains: hemolysin, enterotoxin
- Proteins that mediate plasmid transfer to uninfected strains
Conjugative Plasmid transfer
- Some plasmids have specific transfer genes, can copy DNA, transfer one copy to a plasmid- cell
- Sex pilus: rigid fiber, sticks out from cell wall. Acts as recognition molecule to locate sensitive cell (plasmid minus). When + and - cell are attached by pilus, cells pulled together. (Grappling hook analogy).
- Plasmid DNA replicates during transfer; one copy remains in donor cell, identical copy transferred to recipient.
- Once recipient receives plasmid, it grows pili, becomes a donor. One plasmid could potentially infect a whole population, convert all cells to plasmid-containing cells.

F-factors: F+, F-, and Hfr strains
- F is a conjugative plasmid. Normally, F+ x F- ----> all progeny F+
- Very rarely, F plasmid becomes integrated into host DNA = Hfr. Very stable, doesn't "pop" back out
- Still retains ability to transfer to another cell, but in doing so, drags host DNA with it
- Transfer is always unidirectional; always moves from plasmid + cell to plasmid - cell
- Transfer always originates at same point (due to location of F factor in chromosome at one site)
- Transfer occurs with fairly high frequency (Hfr = abbreviation).
Mating of Hfr with F-
- In E. coli, when Hfr mates with F-, transfer of entire chromosome takes ~100 minutes. Location of different genes can be mapped by time of transfer.

- To detect transfer, must design donor and recipient cells cleverly.
- Donor cell: must have easily selectable markers (typically, carries wild-type genes that allow growth). But must somehow prevent donor cells from growing after DNA transfer. Trick: make donor sensitive to some antibiotic (e.g., streptomycin sensitive), add that antibiotic to plates to look for successful gene transfer. Result: donor cells all killed on plates.
- Recipient cell: must carry marker genes to identify portions of map. Typically use auxotroph markers (e.g., his-, leu-, thr-, ser-, etc.), plate cells onto plates lacking one needed metabolite (e.g. histidine) but containing all other metabolites. Result: none of "native" recipient cells alone can grow.
- Overall result: Only cells that will be able to grow are recipient cells that received a wild type gene by transfer, can grow in absence of missing metabolite.
- Note: in practice, must plate cells at different time points on a variety of different plates, all containing antibiotic: one set of plates lacking histidine, another set lacking leucine, still another set lacking threonine, etc.
- Look at # of colonies growing on each plate, plot results vs. time. See graph. Extrapolate curve for each marker back to earliest possible time of entry.
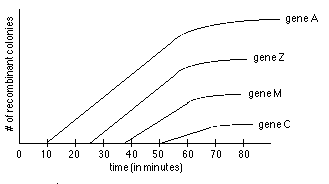
- In example above, can determine two things:
- gene order is A Z M C on DNA (not A C M Z or something else)
- relative location of genes along map is about as follows: A at 10 min, Z at 26 min, M at 37 min, C at 51 minutes along map
Linear and Circular genetic maps
- Is temporal map same as physical piece of DNA? Yes, many experiments support this.
- Is DNA really linear? No, physical evidence suggests a circle. In fact, by using different types of Hfr donors (with point of F-factor insertion at different points on DNA), can get different linear maps; when comparing these, circular map is logically necessary
- Linear map will always result from using a particular Hfr strain, since DNA must be transferred as linear molecule.
Applications of conjugation mapping
- Can use conjugation to find approximate location of any new gene. Note: E. coli has ~ 3000-5000 genes (predicted); about 1/2 of these are currently known. Still many to discover. Valuable tool to help identify features of a gene.
- For fine mapping, can use transduction (more sensitive for small differences between genes close together), eventually use physical methods (restriction fragment mapping, DNA sequencing).
- Most bacteria don't have characterized conjugation systems; seriously limits how much progress we can make in trying to discover and arrange genes in these microbes.
Take a Self-Quiz on this material
Return to Lecture Index
Return to MCB 229 Course Resources page