II. Reactions between Delta-Aminolevulinic Acid (ALA) and Protoporphyrin IX (Proto)
A. Biosynthesis of ALA
B. Biosynthesis of Porphobilinogen (PBG)
C. Biosynthesis of Uroporphyrinogen III (Urogen III)
D. Biosynthesis of Coproporphyrinogen III (Coprogen III)
E. Biosynthesis of Protoporphyrinogen IX (Protogen IX)
F. Biosynthesis of Protoporphyrin IX (Proto IX)
G. References
II. Reactions Between Delta-Aminolevulinic Acid (ALA) and Protoporphyrin IX (Proto)
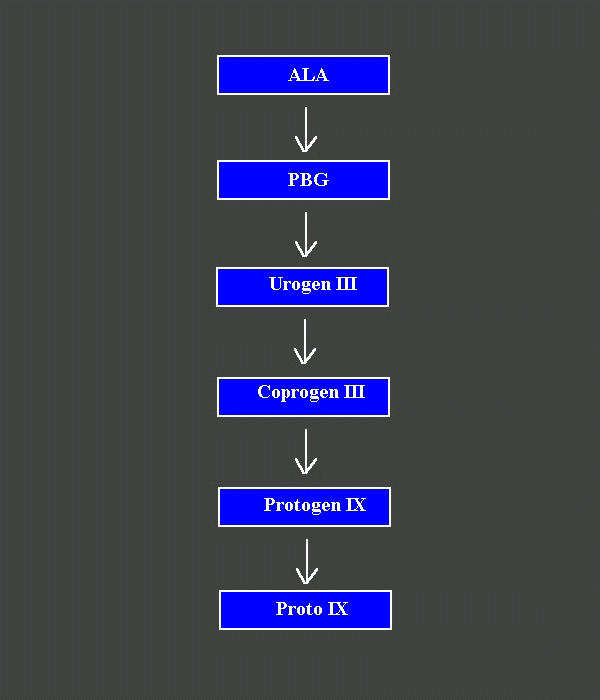
Fig. 1. The sequence of biosynthetic reactions between ALA and Proto.
A. Biosynthesis of Delta-Aminolevulinic Acid (ALA)
Delta-aminolevulinic acid (5-aminolevulinic acid) is the building block of all tetrapyrroles in nature. It is synthesized via a different pathway in animal cells and lower plants than in green plants.
1. In Animal Cells and Lower Plants
In animal cells and lower plants, ALA is formed by condensation of glycine and succinyl-CoA (Gibson et al , 1958) . The reaction is catalyzed by ALA synthetase and takes place in the mitochondria. The biosynthesis of succinyl-CoA from succinic acid is catalyzed by succinyl-thiokinase in the presence of Mg ++ and ATP, and takes place also in the mitochondria (Gibson et al , 1958). ALA is exported to the cytoplasm for further metabolism (Granick, 1963).
2. In Green Plants
In green plants ALA is formed from glutamic acid (Beale and Castelfranco, 1974) via three reactions (Kannangara et al, 1984). In a first reaction, glutamate- tRNAGlu ligase catalyzes the ligation of glutamate to tRNA in the presence of ATP and Mg++. In a second reaction, the glutamyl-tRNA complex is converted into a linear glutamate semialdehyde (GSA) by NADPH:Glu-tRNA(oxido)reductase (also called glutamyl-tRNA dehydrogenase) (Kannangara et al, 1984) or into a cyclic GSA (hydoroxyaminotetrahydropyranone, HAT for short) (Jordan et al, 1989). Finally, GSA aminotransferase converts GSA to ALA in the presence of vitamin B6 and pyridoxal phosphate. These reactions take place in the stroma of the plastid.
B. Biosynthesis of Porphobilinogen (PBG)

PBG is formed from two molecules of ALA and liberates two molecules of water. The dimerization reaction is catalyzed by ALA dehydratase also known as PBG synthase (Gibson et al, 1955; Schmid and Shemin, 1955). The enzyme binding sites of the two ALA substrates have been designated the A (gives rise to the acetic side chain of PBG) and P (gives rise to the propionic side chain of PBG) sites. The first substrate binds to the P site where it forms a Schiff base with the enzyme. The second ALA molecule interacts with the A site (Jordan and Seehra, 1980) to form an enzyme-two substrate complex. The precise mechanism by which the 5-membered PBG ring is formed from the enzyme-two substrate complex is till uncertain. In animal cells PBG is formed from ALA in the cytoplasm (Rebeiz et al, 1996). In plants, PBG synthase is loosely bound to the plastid membranes (Lee et al, 1991).
C. Biosynthesis of Uroporphyrinogen III (Urogen III)

Urogen III is the universal precursor of all metabolic tetrapyrroles (Neve and Labbe, 1956). Its biosynthesis from PBG requires the cooperation of two enzymes, PBG deaminase (Bogorad, 1958) and Urogen III synthase also known as cosynthetase (Bogorad, 1958). In E. coli PBG deaminase is coded for by the gene hemC (Thomas and Jordan, 1986). The apoprotein consists of 353 amino acids with a molecular weight of 34245. The active site contains two constitutive PBG molecules (dipyrromethane cofactor) attached to the apoprotein by a cysteine residue (Cys-242) (Hart et al, 1987; Jordan and Warren, 1987). In a first step, one PBG molecule binds to the deaminase. A covalent bond is formed between the second constitutive PBG molecule and the first PBG substrate, and one molecule of ammonia is released. This first condensation leads to the formation of ring A of Urogen III. This step is repeated three more times and results in the formation of an open chain tetrapyrrole which is displaced from the enzyme by water to yield 1-hydroxymethylbilane (HMBL) (also called preuroporphyrinogen) (Battersby et al, 1979; Jordan and Seehra, 1979; Battersby et al, 1983; Battersby et al, 1982). Hydroxymethylbilane is unstable and in the absence of the cosynthetase cyclases at neutral pH to yield Urogen I. In the presence of the cosynthetase, hydroxymethylbilane is rapidly converted into Urogen III (Battersby et al, 1982b). This reaction involves inversion of ring D of HMBL and cyclization with the release of one water molecule. In E. coli the cosynthetase is coded for by the gene hemD (Jordan et al, 1988). The apoprotein consists of 246 amino acids with a molecular weight of 27766. HemC and hemD occur in tandem and overlap by one base pair. In animal cells, Urogen III is formed in the cytoplasm (Rebeiz et al, 1996). In plant cells, PGB deaminase and the cosynthetase are loosely bound to the plastid membranes (Lee et al, 1991)
D. Biosynthesis of Coproporphyrinogen III (Coprogen III)
| |

Urogen III is the branching point where the biosynthesis of vitamin B12 diverges from that of heme and Chl. Urogen decarboxylase converts Urogen III to Coprogen III (Granick and Mauzerall, 1958; Mauzerall and Granick, 1958). Stepwise decarboxylation of the 4 acetate side chains and the resulting structures of the intermediates led to the proposal that the acetate side chains on rings D, A, B. and C are decarboxylated in a clockwise fashion starting with ring D(Jackson et al, 1976; JJackson et al, 1980). Although this appears to be the case in patients suffering from porphyria cutanea tarda, a random rather than an ordered decarboxylation appears to prevail in normal individuals (Luo and Lim, 1993). This led to the proposal that the substrate binding site has such a flexible architecture that at low Urogen concentrations, decarboxylation may be ordered, while at high substrate concentrations it may be random (Akhtar, 1984). The DNA coding for Urogen III decarboxylase in humans (Romeo et al, 1986) and rats (Romana et al, 1987) has been cloned and sequenced. The human enzyme consists of 367 amino acids with a molecular weight of 40,831. In animal cells Coprogen III is formed in the cytoplasm (Rebeiz, et al, 1996). In plants Urogen III decarboxylase appears to be loosely bound to the plastid membranes (Lee et al, 1991)
E. Biosynthesis of Protoporphyrinogen IX (Protogen IX)

Conversion of Coprogen III to Protogen III involves oxidative decarboxylation of the two propionate side chains on rings A and B and their conversion to vinyl (Sano and Granick, 1961). The mammalian enzyme has
an absolute requirement for oxygen, but requires no reducing agent. Recent studies indicate that the mammalian enzyme is a dimer of two 37000 subunits (Kohno et al, 1993). The observation of harderopoprphyrinogen accumulation (one vinyl at position 2 and one propionate at position 4)(Sano and Granick, 1961) and its subsequent isolation (Kennedy et al, 1970) led to the proposal that the decarboxylation of ring A occurs before that of ring B. The precise mechanism of oxidative decarboxylation is still uncertain. In animal cells cytoplasmic Coprogen III is transported to the mitochondria in an ATP-dependent process where it is converted to Protogen IX (Rebeiz, et al, 1996). In plant cells, Coproporphyrinogen oxidase appears to be loosely bound to the plastid membranes (Lee et al, 1991).
F. Biosynthesis of Protoporphyrin IX (Proto IX)

Protoporphyrinogen IX oxidase (Protox for short) catalyzes the conversion of Protogen IX to Proto (Poulson and Polglase, 1975, Jacobs, and Jacobs, 1982). The oxidation involves the removal of 4 peripheral (meso) hydrogens and two inner hydrogens from the pyrrole nitrogens. In aerobic organisms, oxygen is the oxidant. Removal of the hydrogens appear to be stereospecific (Battersby, et al, 1976). The enzyme has been purified to apparent homogeneity from bovine liver (Siepker et al, 1987). It appears to be a monomer with a molecular weight of approximately 65,000. The bovine enzyme has a tightly bound FAD prosthetic group. The plant enzyme has been partially characterized (Jacobs and Jacobs, 1987). Diphenyl ether herbicides inhibit Protox (Matringe et al, 1989) and result in the accumulation of Protogen IX which translocates to various part of the cell (Lehnen et al, 1990). When Protogen is converted to Proto at various cellular sites, cell death ensues in the light in a typical porphyrin-dependent photodynamic herbicidal phenomenon (Rebeiz, et al, 1984). In animal cells, during heme biosynthesis, Protogen IX is converted to Proto IX in the mitochondria. In plant cells, Protox appears to be loosely bound to the plastid membranes (Lee et al, 1991).
G. References
- Gibson, H. D., Laver, W. G., and Neuberger, A. (1958). Initial stages in the biosynthesis of porphyrins. II. The formation of 5-aminolevulinic acid from glycine and succinyl-CoA by particles from chicken erythrocytes. Biochem. J. 70:71-81.
- Granick, S. (1963). The pigments of the biosynthetic chain of chlorophyll and their interactions with light. In Proceedings of the Fifth International Congress of Biochemistry Vol VI, pp 176-186, Pergamon Press, New York.
- Beale, S. I., and Castelfranco, P. A. (1974). The biosynthesis of delta-aminolevulinic acid in higher plants. II. Formation of 14C-delta-aminolevulinic acid from labelled precursors in greening plant tissues. Plant Physiol. 53: 297-296.
- Kannangara C. G., Gough, S. P., Oliver, R. P., and Rasmussen, S. K. (1984). Biosynthesis of delta-aminolevulinic acid in greening barley leaves. VI. Activation of glutamate by ligation to RNA. Carlsberg Res. Commun. 49:417-437.
- Jordan, P. M., Cheung, R. P., Sharma, R. P., and Warren, M. J. (1989). A cyclic intermediate, 2-hydroxy-3-aminotetrahydropyran-1-one (HAT) as a precursor for 5-aminolevulinic acid in greening barley, Tet. Lett. 34:1177-.
- Gibson, K. D., Neuberger, A., and Scott, J. J. (1955). The purification and properties of 5-aminolevulinic acid dehydratase. Biochem. J. 70: 618-629.
- Schmid, R. and Shemin, D. (1955). The enzymic formation of porphobilinogen from 5-aminolevulinic acid and its conversion to protoporphyrin. J. Am. Chem. Soc.77: 506-508.
- Jordan, P.. M., and Seehra, J. S. (1980). 13C NMR as a probe for the study of enzyme catalyzed reactions. Mechanism of action of 5-aminolevulinic acid dehydratase. FEBS Lett. 114: 283-286.
- Rebeiz, N., Arkins, S., Kelley, K. W., and Rebeiz, C. A. (1996). Enhancement of coproporphyrinogen III Transport into isolated transformed leukocyte mitochondria by ATP. Arch. Biochem. Biophys. 333: 475-481.
- Lee, H. J., Ball, M. D., and Rebeiz, C. A. (1991). Intraplastidic localization of the enzymes that convert d-aminolevulinic acid to protoporphyrin IX in etiolated cucumber coryledons. Plant Physiol. 96: 910-915.
- Neve, R. A., and Labbe, R. F. (1956). Reduced Uroporphyrinogen III in the biosynthesis of heme. J. Am. Chem. Soc. 78: 691-692.
- Bogorad, L. (1958). The enzymic synthesis of porphyrins from porphobilinogen. I. Uroporphyrinogen I. J. Biol. Chem. 233: 501-509.
- Bogorad, L. (1958). The enzymic synthesis of porphyrins from porphobilinogen. II. Uroporphyrin III. J. Biol. Chem. 233: 510-515.
- Thomas, S. D. and Jordan, P. M. (1986). Nucleotide sequence of the hemC locus encoding porphobilinogen deaminase of Escherichia coli K12. Nucleic Acids Res. 14: 6215-6226.
- Hart, G. J., Miller, A. D., Leeper, F. J., and Battersby, A. R. (1987). Biosynthesis of the natural porphyrins: proof that hydroxymethylbilane synthase (porphobilinogen deaminase) uses a novel binding group in its catalytic action. J. Am. Chem. Soc. Chem. Commun..1762-1765.
- Jordan, P. M., and Warren, M. J. (1987). Evidence for a dipyrromethane cofactor at the catalytic site of E. coli porphobilinogen deaminase. FEBS Lett. 225: 87-92.
- Battersby. A. R., Fookes, C. J. R., Matcham, G. W. J. and Mc.Donald, E. (1979). Order of assembly fo the four pyrrole rings during the biosynthesis of the natural porphyrins. J. Chem. Soc. Chem. Commun..539-541.
- Jordan, P. M. and Seehra, J. S. (1979). The biosynthesis of uroporphyrinogen III: order of assembly of the four porphobilinogen molecules in the formation of the tetrapyrrole ring. FEBS Lett. 104: 364-366.
- Battersby. A. R., Fookes, C. J. R., Matcham, G. W. J., Mc.Donald, E., and Hollenstein, R. (1983). biosynthesis of porphyrins and related macrocycles. Part 20. Purification of deaminase and studies on its mode of action. J. Chem. Soc. Perkins Trans. 1: 3031-.
- Battersby. A. R., Fookes, C. J. R., Gustafson-Potter, K. E., Mc.Donald, E. and Matcham, G. W. J. (1982a). Biosynthesis of porphyrins and related macrocycles. XVII. Chemical and enzymic transformation of isomeric aminoethylbilanes into uroporphyrinogens: proof that unrearranged bilnae is the preferred enzymic substrate and detection of a transient intermediate. J. Chem. Perkin Trans. 1:2413-
- Battersby. A. R., Fookes, C. J. R., Gustafson-Potter, K. E., Mc.Donald, E. and Matcham, G. W. J. (1982b). Biosynthesis of porphyrins and related macrocycles. Part 18. Proof by spectroscopy and synthesis that unarranged hydroxybilane is the product from deaminase and the substrate for cosynthetase in the biosynthesis of uroporphyrinogen-III. J. Chem. Soc. Perkins Trans. 1: 2427-
- Jordan, P. M., Mgbeje, B. I. A., Thomas, S. D., and Alwan, A. F. (1988). Nucleotide sequence of the hemD gene of Escherichia coli encoding uroporphyrinogen III synthetase and initial evidence for a hem operon. Biochem. J. 249: 613-616.
- Granicks, and Mauzerall, D. (1958). Porphyrin biosynthesis in erythrocytes. II. Enzymes converting delta-aminolevulinic acid to coproporphyrinogen. J. Biol. Chem. 232:1119-1140.
- Mauzerall, D. and Granicks, S. (1958). Porphyrin biosynthesis in erythrocytes. III. Uroporphyrinogen and its decarboxylation. J. Biol. Chem. 232:1141-1162.
- Jackson, A. H., Sancovich H. A., and Ferramola, A. M., Evans N., Games, D. E., and Matlin, S. E. (1976). Macrocyclic intermediates in the biosynthesis porphyrins. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 273:191-206.
- Jackson, A. H., Sancovich H. A., and Ferramola, A. M. (1980). Synthetic and biosynthetic studies of porphyrins. III. Structures of intermediates between uroporphyrinogen III and coproporphyrinogen III: synthesis of fourteen heptacarboxylic, hexacarboxylic and pentacarboxylic porphyrins related to uroporphyrin III. Bioorg. Chem. 9: 71-120.
- Luo J., and Lim, C. K. (1993). Order of uroporphyrinogen III decarboxylation on incubation of porphobilinogen and uroporphyrinogen III with erythrocyte uroporphyrinogen decarboxylase. Biochem. J. 289: 529-532.
- Akhtar, M. (1994). The modification of acetate and propionate side chains during the biosynthesis of haem and chlorophylls: mechanistic and stereochemical studies. In: The Biosynthesis oh Tetrapyrrole Pigments . Ciba Foundation Symposium, 180. pp 131-155, John Wiley and Sons, Chichester.
- Romeo, P. H., Raich, N., Duhart, A., Beaupain, D., Pryor, M., Kushner, J., Cohen-Solal, M., and Goossens, M. (1986). Molecular cloning and nucleotide sequence of a complete human uroporphyrinogen decarbozylase cDNA. J. Biol. Chem. 261: 9825-9831.
- Romana, M., LeBoulch, P. and Romeo, P. H. (1987). Rat uroporphyrinogen decarboxylase cDNA: nucleotide sequence and comparison to human uroporphyrinogen decarboxylase. Nucleic Acids Res. 15:7211.
- Sano, S. and Granick, S. (1961). Mitochondrial coproporphyrinogen oxidase and protoporphyrin formation. J. Bol. Chem. 236:1173-1180.
- Kohno, H., Furukawa T., Yoshihaga, T., Tukunaga, R., and Taketani, S. (1993). Coproporphyrinogen oxidase. J. Biol. Chem. 268:21359-21363.
- Kennedy, G. Y., Jackson, A. H., Kenner, G. W., and Suckling, C. J. (1970). Isolation, structure and synthesis of a tricarboxylic porphyrin from harderian glands of rat. FEBS Lett. 7: 205-206.
- Poulson, R., and Polglase, W. J. (1975). The enzymic conversion of protoporphyrinogen IX to protoporphyrin IX. Protoporphyrinogen oxidase activity in mitochondrial extracts of Saccharomyces cerevisiae. J. Biol. Chem. 250:1269-1274.
- Jacobs, N. J., and Jacobs, J. M. (1984). Protoporphyrinogen oxidation, an enzymatic step in heme and chlorophyll synthesis: Partial characterization of the reaction in plant organelles and comparison with mammalian and bacterial systems. Arch. Biochem. Biophys. 229:312-319.
- Battersby, A. R., Mc. Donald, E., Redfern, J. R., Staunton, J. and Wightman, R. H. (1976). Biosynthesis of porphyrins and related macrocycles. V. Structural integrity of the type III porphyrinogen macrocycle in an active biological system; studies on the aromitazation of protoprphyrin-IX. J. Chem. Soc. Perkins Trans. I: 266-273.
- Siepker, L. J., Ford, M., de Kock, R., and Kramer, S. (1987). Purification of bovine protoporphyrinogen oxidase: immunological cross-reactivity and structural relationship to ferrochelatase. Biochim. Biophys. Acta. 913: 349-358.
- Jacobs, J. M., and Jacobs, N. J. (1987). Oxidation of protoporphyrinogen to protoporphyrin, a step in the in chlorophyll and heme biosynthesis: purification and partial characterization of the enzyme from barley organelles. Biochem. J. 244:219-224.
- Matringe, M., Camadro, J. M., Labbe, P., and Scalla, R. (1989). Protoporphyrinogen oxidase as a molecular target for diphenyl ether herbicides. Biochem. J. 260: 231-235.
- Lehnen, L. P. Jr., Sherman, T. D., Beceril, J. M., and Duke, S. O. (1990). Tissue and cellular localization of acifluorfen-induced porphyrins in cucumber cotyledons. Pest. Biochem. Physiol. 37:239-248.
- Rebeiz, C. A., Montazer-Zouhoor, A., Hopen, H. J. and Wu, S. M. (1984). Photodynamic herbicides: Concept and phenomenology. Enzyme and Microbial Technology 6:390-401.
Go Back to Main Menu